About Database
Living organisms exhibit diverse forms of DNA distinct from B-DNA, each serving unique biological functions. The conformation adopted by a DNA molecule depends primarily on factors such as hydration level, DNA sequence, chemical modifications of bases, and the types and concentrations of metal ions in the solution. The seven principal non-B DNA conformations include A-phased repeats, direct repeats and slipped motifs, G-quadruplex forming motifs, inverted repeat and cruciform motifs, mirror repeats and triplex motifs, Z-DNA motifs, and short tandem repeats.
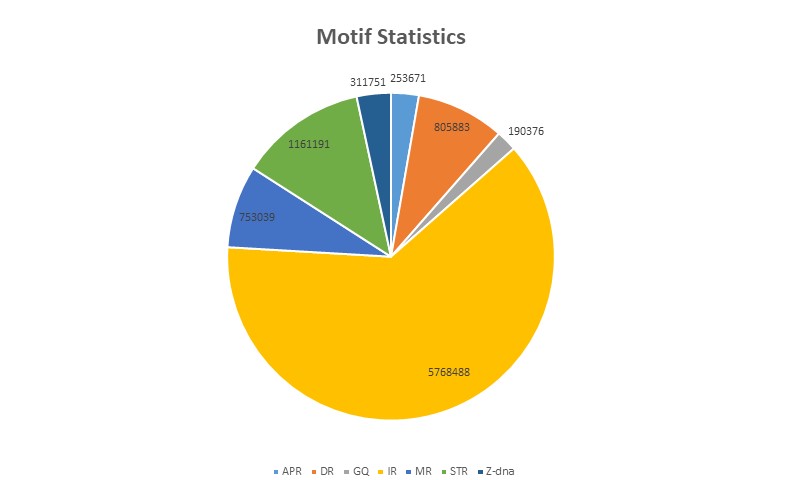
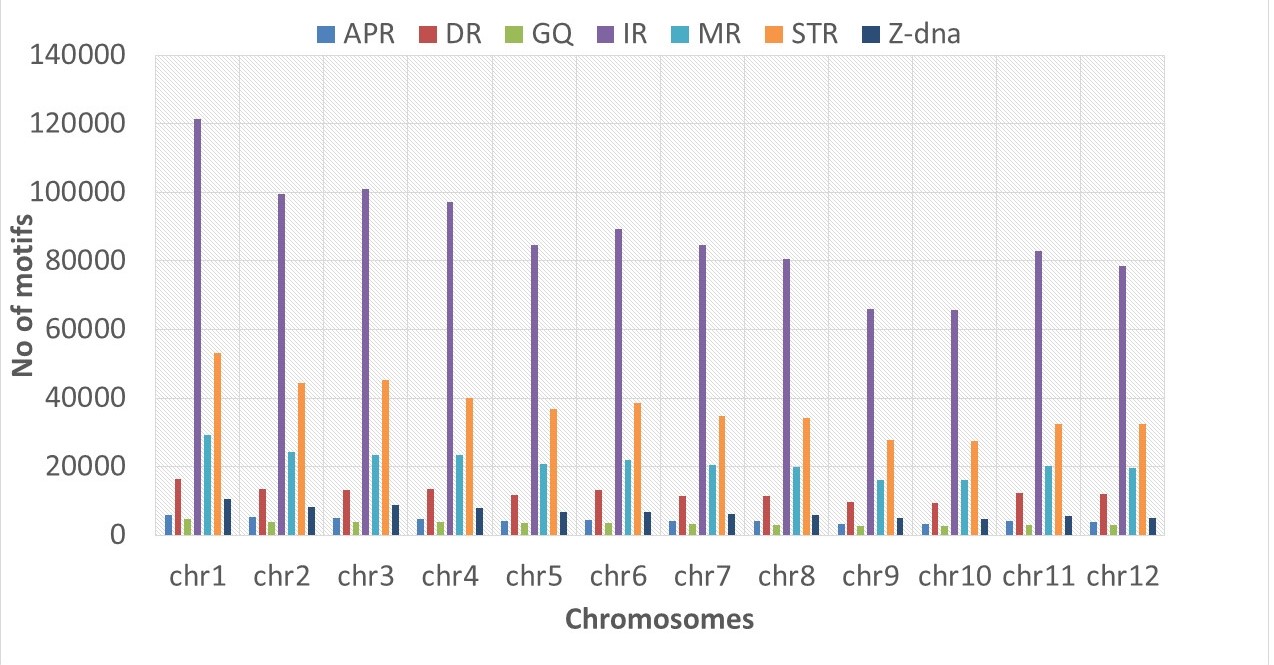
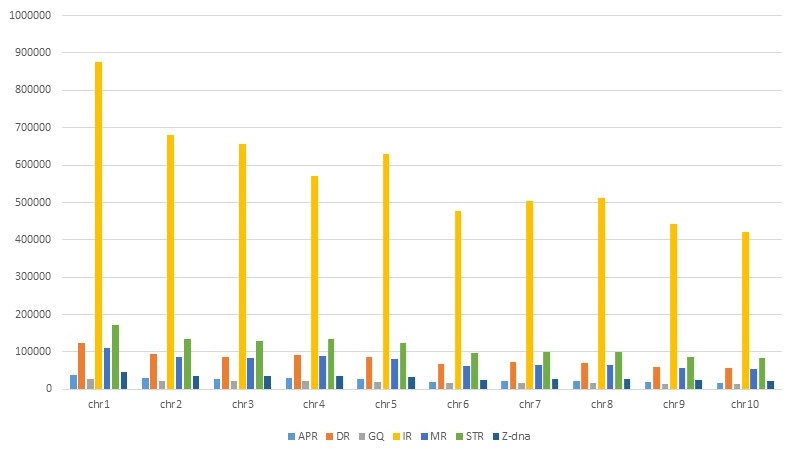
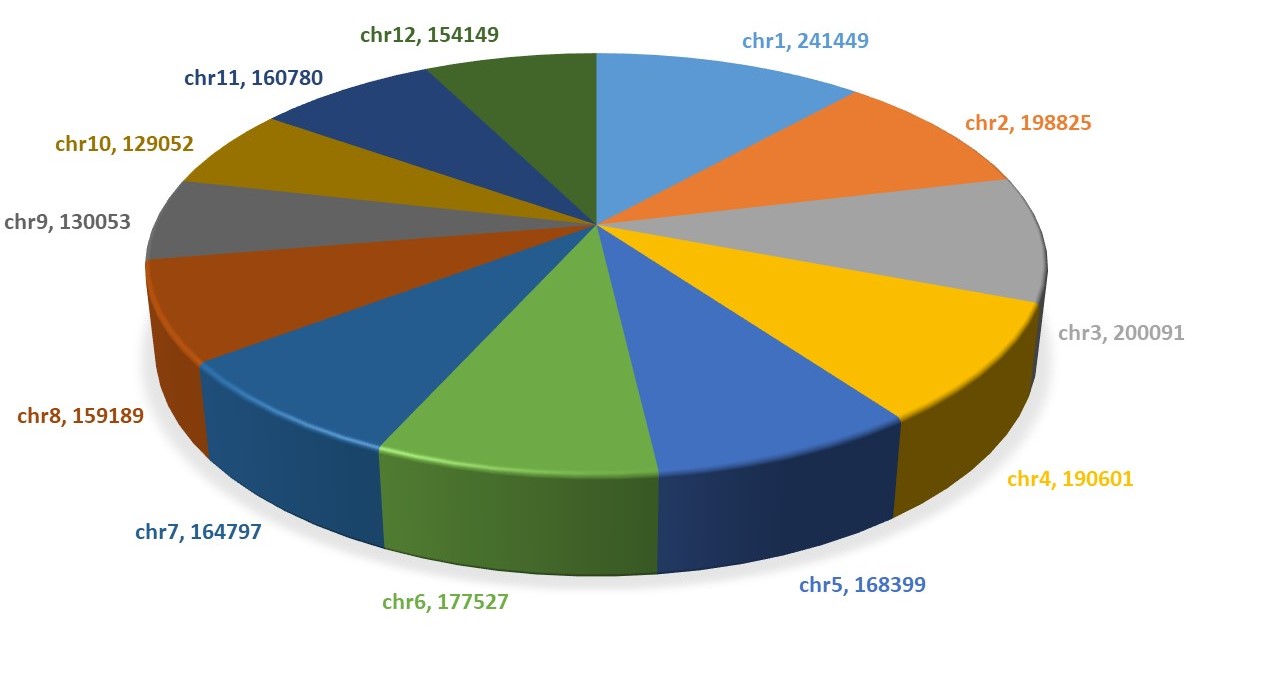
In the Rice Genome, the numbers of A-phased repeats, direct repeats, G-quadruplex motifs, inverted repeats, mirror repeats, short tandem repeats, and Z-DNA motifs are 2, 7, 2, 51, 12, 22, and 4, respectively. Similarly, in the Maize Genome, these numbers are 3, 9, 2, 62, 8, 13, and 3, respectively. The Non B-DNA database encompasses seven major non-B DNA motifs identified in rice and maize crops, providing additional information about the annotation of genes harboring these motifs.